3rd Annual University of Auckland Mass Spectrometry Symposium
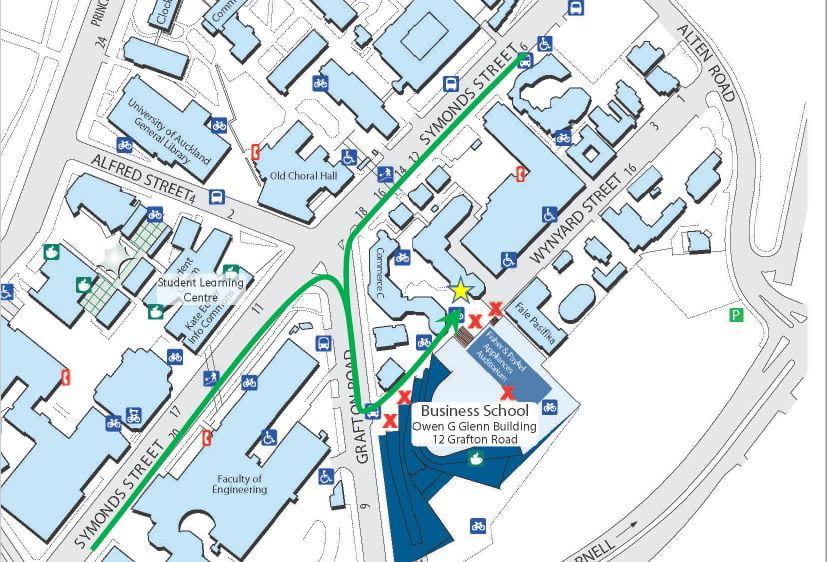
CIty Campus (Arts), UoAPlease join us for the 3rd Annual University of Auckland Mass Spectrometry Symposium. This year our symposium will showcase a diverse array of applied and fundamental mass spectrometry research from overseas and across the university.
Keynote speakers Assistant Professor Tiffany Porta Siegel (Maastricht MultiModal Molecular Imaging (M4I) institute) will describe the use of mass spectrometry imaging in clinical research and developments in ambient ionisation MS for intra-operative applications, while Dr Farhana Pinu (Plant and Food Research) will present her research on the application of mass spectrometry to systems biology in grape and wine analysis. Other researchers will contribute to a day of talks covering areas of mass spectrometry such as metabolomics, proteomics, clinical analysis and volatilomics. There will also be talks dedicated to mass spectrometry data analysis.
We acknowledge the MaSH, the School of Medical Sciences, the Faculty of Science and the Liggins Institute for supporting this event. Please RSVP to Martin Middleditch for catering purposes. Draft program and abstracts below…
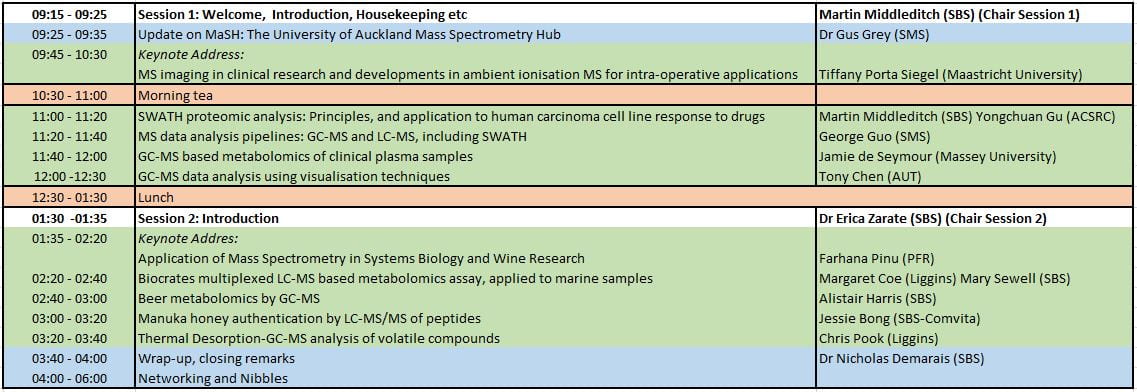
Draft Program:



Symposium Abstracts:
George Guo
Playing with LEGO? Build a MS data processing pipeline at your service.
Abstract: Find mass spectrometry data processing quite complicated? Welcome to the MS data playground. Here, you will learn the generic MS data formats and hundreds of useful modules for your data processing purpose. All you need to do is put them together and wait for the magic. George’s talk will lead you through each steps from spectrum and chromatogram peak picking to statistical features analysis.
Tony Chen
Computer vision in GCMS, a machine learning approach to untargeted GCMS analysis
Abstract: Computer vision has been very successful in many areas like face recognition, object detection, image classification, this talk will describe how to use a convolution neural network to achieve sample classification on raw
Yongchuan Gu
Quantifying the protein expressions at the whole proteome level – Are we there yet?
Abstract: Over the past decade, gene expression at the whole genome level has been routinely accessible using RNA-based methods, but proteomics is rapidly coming of age because of major advances in mass spectrometry (MS). To fully identify and quantify the entire complement of proteins in biological samples of interest has long been the ‘Holy Grail’ of proteomics. Data-Independent Acquisition (DIA) is a relatively new MS-based technique for unbiased systematically collecting MS/MS data, which allows comprehensively measuring every peptide in a protein digest. It also gives flexibility (e.g. retrospective data analysis) to investigate multiple hypotheses without having to acquire additional data. In this talk, I will describe how the technology is being developed in the ACSRC with the help from University Mass Spectrometry Hub (MaSH), and its application in identifying changes in the proteome of cancer cells under hypoxia.
Jamie de Seymour
Can we find diet-associated biomarkers of cognition in the plasma of older adults using mass spectrometry?
Abstract: By 2051, a quarter of the New Zealand population will be over 65 years of age; finding ways to improve cognitive function in older adults will help to delay cognitive decline and enable them to live independently for longer. The aim of our project is to investigate the plasma metabolome of older independent-living adults (65-74 years) for biomarkers of cognitive function. Furthermore, we plan to investigate whether any of the cognition-associated biomarkers are linked with dietary intake.
Farhana Pinu
Application of Mass Spectrometry in Microbial Systems Biology and Wine Research
Abstract: Here, I will present quantitative mass spectrometry (MS) based metabolomics data that show the seasonal and geographical differences in New Zealand Sauvignon blanc grapes and wines and how these datasets are being used to develop system biology platforms for a commercial wine yeast strain.
Margaret Coe
Metabolomics the fast and easy way
Abstract: Our lab recently used the Biocrates MetIDQ kit for quantitative and qualitative analysis of 407 analytes comprised of amino acids, biogenic amines and lipids. The details about the kit and some of the results produced will be shown.
Alistair Harris
Beer metabolomics by GC-MS”, would you like to change it?
Abstract: Sound is ubiquitous in nature and industry, however it’s impact on microbes is largely unknown. We are using a Saccharomyces cerevisiae strain to investigate the effect audible sound has on beer fermentation and the associated metabolites, with the ultimate aim of producing a novel sonic beer.
Jessie Bong
Peptide profiling as a novel and alternative approach to mānuka (Leptospermum scoparium) honey authentication
Abstract: Proteomics is an emerging tool in food authentication that has not been optimised for honey analysis. We applied a bottom-up proteomics approach using the high-resolution nanoLC-QqTOF-MS/MS system (Sciex Triple TOF 6600) and identified 12 unique peptides markers in mānuka honey that can be useful for the honey fingerprinting. Nectar analyses confirmed the origin and specificity of these peptides to L. scoparium nectar, and application of the method to NZ honey analyses demonstrated promising results in line with other chemical marker analyses.
Chris Pook
Applications of untargeted volatilomics by Thermal Desorption Gas Chromatography with Mass Spectrometry
Thermal Desorption (also Thermal Extraction) is a technique for extracting volatile compounds from samples or sorbents using a flow of inert gas, high temperatures and cryotrapping of the extracted material for analysis by GC-MS. Recent technical advances in the field of TD-GC-MS have led to a rapid growth in its popularity due to the simplicity, robustness and reproducibility of the analysis as well as the greatly improved sensitivity over existing volatile sampling technologies, such as Static Headspace Sampling and Solid Phase Microextraction. This technique has potentiated the emergence of volatilomics: the analysis of all volatile compounds present in a solid, liquid or gaseous volume. This presentation will present some applications of this powerful technique to biological and environmental analysis.
